







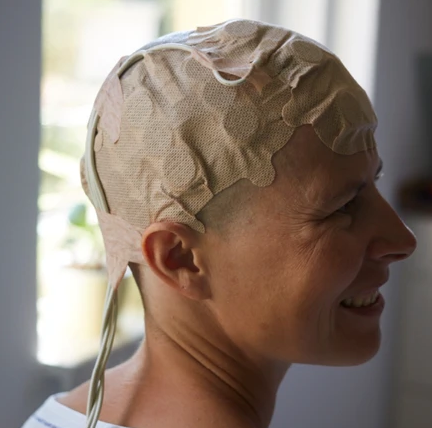
Opting out of Optune
A treatment with small benefit and notable costs leads to a tough call.




Beating Swords into Ploughshares
A returning WWII veteran takes up an unlikely field of study.


Sowing Seeds on Rocky Soil
Tracing my physicist grandfather's origins to an unexpected place.
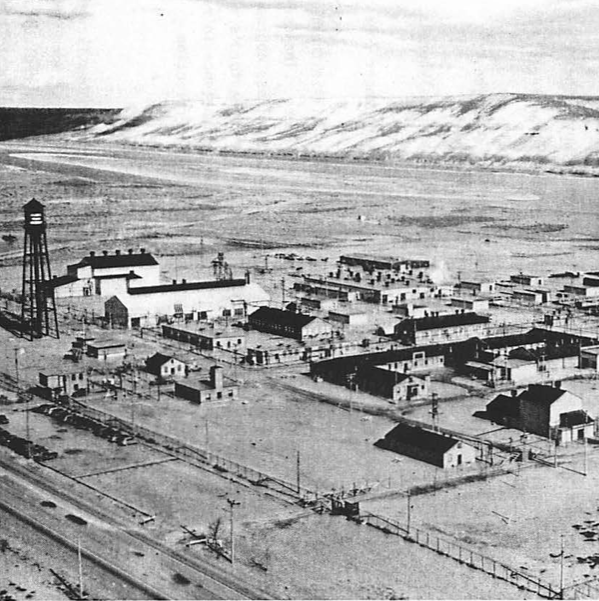

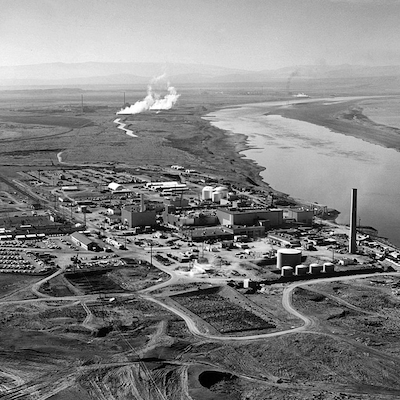
Grandpa, What Did You Do at the Atom Bomb Factory?
First in a series investigating what exactly my grandfather did at the Hanford Engineer works.
How I Bought and Estimated the Purity of Sodium Dichloroacetate
Trust, but verify. On second thought, just verify.

Beware Oncologists Bearing Hope
Why interpretations from this special breed of doctor should be considered carefully.




Just Saying "No" to Adjuvant Temozolomide
Discovering that the medicine cabinet for my terminal disease was filled with fluff.

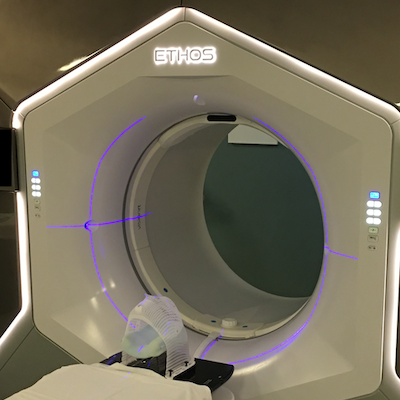

Temodar, MGMT Methylation, and the Endless Loop of Bad Glioblastoma Treatments
Understanding why half of patients will take a drug offering little to no benefit.