A Molecular Language for Modern Chemistry: Reading FlexMol Documents with Octet
An XML language is only as useful as the software tools that take advantage of it. Previous articles have discussed how the XML language FlexMol can solve a variety of molecular representation problems ranging from the multiatom bonding of metallocenes to the axial chirality of biaryls. Octet is a framework written in Java that speaks FlexMol natively. In this article, I'll show how Octet can be used to read a sample FlexMol document.
Prerequisites
For this tutorial, you'll need Ruby Java Bridge (RJB). Previous articles have discussed the installation and use of RJB on Windows and Linux.
A Sample Molecule
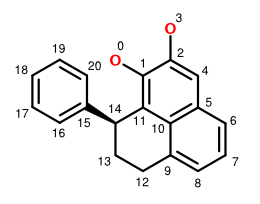
A recent article disused a FlexMol representation of the chiral natural product monolaterol. Using a slightly modified numbering system for this molecule (shown above), we can construct a complete FlexMol representation. In this case, we simply start numbering at index zero, subtracting one from every index in the previous example to match the zero-based indices used in Octet.
A Demonstration Package
To illustrate the process of reading a FlexMol document, I've prepared a small package (demo-20070131.tar.gz) that can be downloaded from SourceForge. In it, you'll find an Octet jarfile (octet-0.8.2.jar), a FlexMol representation of monolaterol (s_monolaterol.xml), a Ruby library (reader.rb), and some Ruby test code (test.rb). Inflate this archive and make it your working directory.
A Simple Test
The following sequence of commands will run the test included with the demonstration package:
export CLASSPATH=./octet-0.8.2.jar
ruby test.rb
You should see several lines of output terminated with the line:
The exact mass of monolaterol is 276.115029755.
You can get more hands-on experience with loading and processing the monolaterol FlexMol document using interactive Ruby (irb). For example:
irb
irb(main):001:0> require 'reader'
=> true
irb(main):002:0> r=Reader.new
=> #<Reader:0x2b9ab173a1f0 @xml_reader=#<#<Class:0x2b9ab1741680>:0x2b9ab1736690>, @handler=#<#<Class:0x2b9ab1741680>:0x2b9ab1736e10>, @builder=#<#<Class:0x2b9ab1741680>:0x2b9ab1736b90>>
irb(main):003:0> mol=r.read_file 's_monolaterol.xml'
=> #<#<Class:0x2b9ab1741680>:0x2b9ab172cd48>
irb(main):004:0> mol.countAtoms
=> 21
irb(main):005:0> mol.countBondingSystems
=> 24
Of course, this is just scratching the surface of what can be done once a FlexMol document has been loaded by Octet.
Conclusions
Octet makes it possible to convert FlexMol documents into Java object representations that can be accessed through Ruby. With an object representation, the possibilities are limitless. Some simple examples have been provided here. Future articles will illustrate more advanced uses.