An Introduction to the ChemDraw CDXML Format
Although it receives relatively little attention in cheminformatics, ChemDraw may be the most important piece of software in chemistry. Every chemist with experience beyond high school will probably have at least heard of it. ChemDraw is the de facto standard for creating graphics for publications and presentations, with every major journal providing settings for authors. It's also one of the formats accepted by the USPTO for patents and patent applications. Over the years, ChemDraw's list of features has grown from chemical structure and reaction drawing to other tasks, including: biopolymer drawing; physical property calculation; name/structure conversion; NMR simulation; and database integration. On ChemDraw's 30th anniversary, C&EN ran a feature about the software's transformative role in chemistry.
ChemDraw stores documents in one of two formats: CDX and CDXML. CDX is a binary format, whereas CDXML is an XML-based file format. With minor differences, both formats share similar semantics. For this reason, this article will refer to the combined binary and XML ChemDraw format as CDX/ML. Although isolated CDX/ML files may be encountered in the wild, it's more common to find them embedded in Microsoft Office or ChemOffice documents. Often, CDX/ML makes its way from these embedded environments to the outside world via the system clipboard.
This article presents a high-level overview of the ChemDraw CDXML format.
CDXML in a Nutshell
As an XML format, CDXML encodes information in three ways:
- Elements. An XML element consists of a named opening tag and a closing tag wrapped in angle brackets (
<and>). Elements may be nested, yielding a tree. The CDXML documentation refers to elements as "Objects." - Attributes. Each element stores a key/value dictionary as a set of named attributes. Most CDXML attributes are optional.
- Element text. In rare cases (e.g., the
Styleobject), an element may contain text data.
The authoritative specification from Perkin Elmer (PE) offers a starting point for understanding CDXML. Elements ("Objects") and attributes ("Properties") are both enumerated and individually documented. The PE specification lists 263 attributes and 38 elements. Although this documentation is mostly complete, several items are missing. For example, details on the Arrow element are not available; its hyperlink redirects to the Perkin Elmer homepage. Several years ago, I wrote an email to technical support noting this problem, but received no response. Those wanting a complete CDXML specification may need to construct one themselves using reverse engineering techniques.
Aside from the official PE specification, not a lot of third party CDXML documentation has been published. Two years ago, a paper describing an open source system capable of reading CDXML files was published (ChemScanner). Although the paper says little about the CDXML format itself, the source code may be helpful as a kind of documentation. In a similar vein, Open Babel supports reading ChemDraw files, but incompletely. Finally, four previous Depth-First articles discuss the CDX binary format:
- A Brief Introduction to the ChemDraw CDX File Format
- Reading and Translating ChemDraw CDX Files with OpenBabel
- Making Sense of the ChemDraw CDX File Format with CDXHexDumper
- Reading (and Rendering) ChemDraw CDX Files in JavaScript
CDX/ML is an odd cheminformatics file format in that is mixes a molecular graph encoding system with visual elements and styling. For example, a given CDX/ML file may contain a chemical structure together with a TLC plate. Each individual bond can be colored, and the text on atom labels can bear custom colors, fonts, and layout instructions.
Sometimes visual elements can carry chemical meaning. For example, an arrow may be part of a reaction scheme. Likewise, a bracket may surround the repeating unit of a polymer.
This broad scope, in which chemically meaningful elements are mixed with visual layout and arbitrary vector graphics, makes CDX/ML one the most complicated file formats in cheminformatics. Coupled with the useful, but incomplete PE specification, CDX/ML is not an easy format to understand or use. The purpose of this article is to fill in some of the blanks.
Objects
Objects (aka "Elements") express the high-level features of every CDXML document. The PE specification enumerates the following Object types:
Document.The root element in the XML tree.Page.A drawing space.Group.An arbitrary collection of Objects.Fragment.Roughly, a molecular graph.Node.Often an atom, but can take many forms.Bond.A connection between twoNodes.Text.A block of styled text runs.Graphic.A geometric shape such as an arc, line, or arrow.Bracketed Group.A group surrounded by brackets.Bracket Attachment.Links the content of a bracketed group with an external feature.Crossing Bond.Links aBracket Attachmentwith aBond.Curve.A Bézier curve.Embedded Object.An operating system-specific object.Table.A grid ofPages.Named Alternative Group.LinksTextwith aGrouporFragment.Template Grid.A grid of templates used by the ChemDraw user interface.Registry Number.Not written or read.Reaction Scheme.An ordered list ofReaction Steps.Reaction Step.A single step reaction consisting of reactants, products, plus signs, arrow(s), reagents, and conditions.Spectrum.Object Tag.A custom object.Sequence.Cross-Reference.Links objects together, possibly across CDXML documents.Border.The edges of a rectangular object, typicallyPage.Geometry.A geometrical relationship between objects.Constraint.A geometrical constraint (distance, angle) between objects.TLC Plate.A thin-layer chromatography plate.TLC Lane.A column within a TLC plate.TLC Spot.A spot on a TLC plate.Splitter.Not used or written.Chemical Property.A property, typically computed by ChemDraw.Color Table.An ordered list ofColors.Color.An RGB color.Font Table.An ordered list ofFonts.Font.A font consisting of name and character set.Style.A run of styled text.Represents Property.Notification that a graphical object represents a chemical property. For example, a circle-plus may represent the formal charge of aNode.Arrow.A graphical arrow with heads, curvature, and other styling.
The most obvious challenge here is the quantity of Objects. As previously noted, ChemDraw combines elements of chemical graphs and vector graphics within the same format. A less obvious challenge is how some Objects seem to recapitulate others (e.g., Geometry vs. Constraint, and Graphic vs. Arrow). ChemDraw is a format that has undergone decades of evolution, a fact evidenced throughout the PE specification.
But perhaps the biggest problem is grappling with the complex ways that the many CDXML Objects can interact with each other.
Entity-Relationship Diagrams
The PE specification provides a lot of the information needed to piece together the relationships between Objects. Unfortunately, the technique used by PE (hyperlinking) doesn't lend itself well to high-level thinking or visualization. What's needed is a way to see the big picture all at once, visually.
Enter Entity-Relationship Diagrams (ERDs). An ERD is a drawing in which the nouns of a system are connected by lines whose terminators represent allowed quantities. An ERD creates a high-level overview of a complex system that can be interpreted and improved by subject matter experts regardless of their level of technical expertise.
The most distinctive feature of an ERD is the graphical expression of quantifier relationships. Quantifiers appear at line terminals, and give information about the cardinality of a relationship. For example, a line terminating with a "zero or one" relationship says that the noun at the end of the line is associated with the noun at the beginning of the line zero or more times. A line without a quantifier indicates no association.
| Quantifier | Meaning |
|---|---|
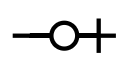 | zero or one |
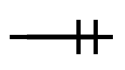 | one and only one |
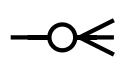 | zero or many |
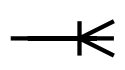 | one or many |
Many drawing tools support the creation and editing of ERDs. One I've found especially useful is draw.io. This open source drawing tool runs in a browser and requires no subscription. ERDs can be stored locally in a custom XML format for later use. Export to SVG and PNG is also supported. Although commercial tools for this kind of thing exist, I find draw.io to be superior in every way. As an added bonus, a VS Code plugin for draw.io is available, allowing the viewing and editing of ERDs from within a project repository.
ChemDraw ERD
Using information presented in the PE CDXML specification, it's possible to construct the following ERD:
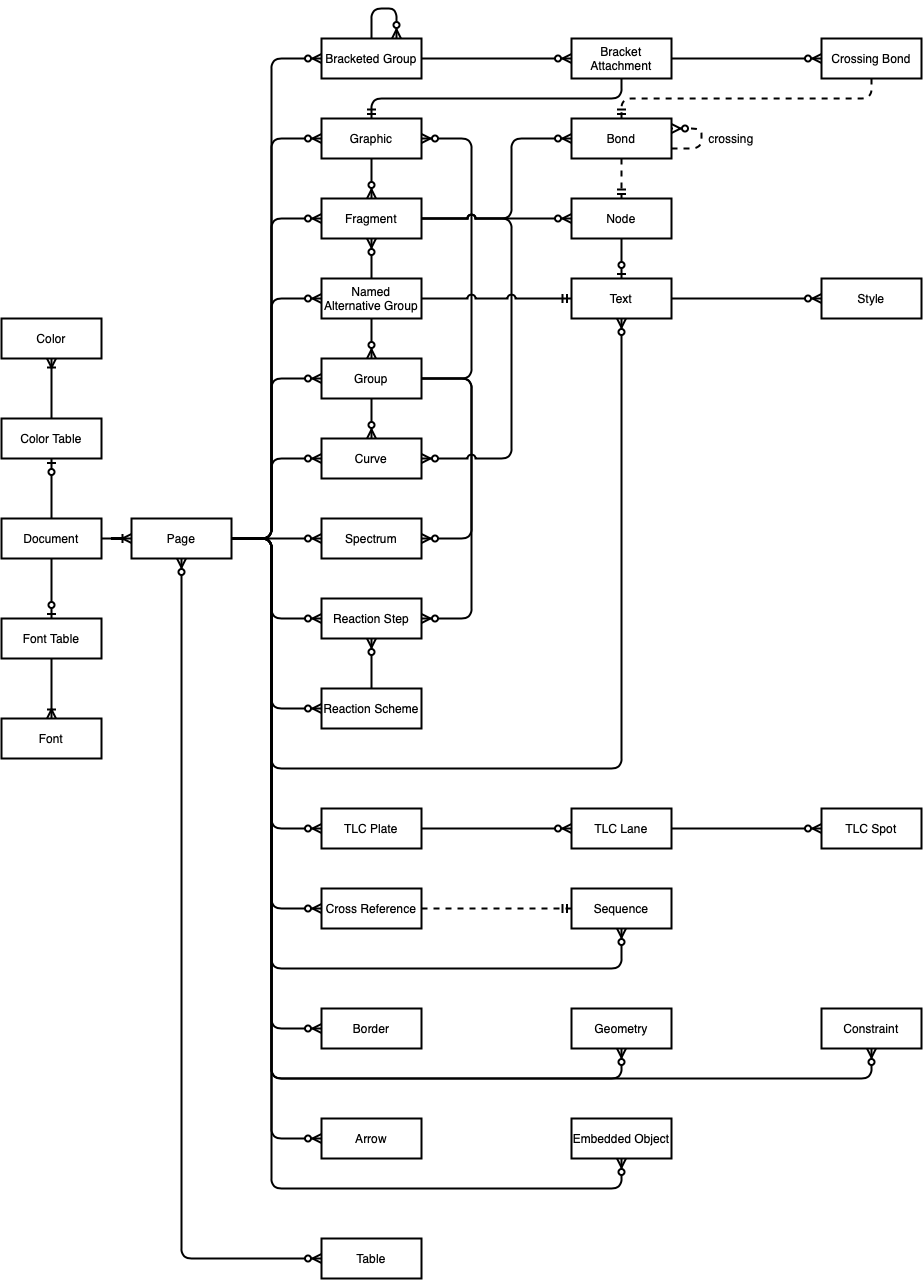
The ERD I created, incomplete though it is, clearly has a lot going on. In fact, there's far too much to absorb non-graphically in my view. The most obvious feature is the large number of Objects associated with a Page. Also apparent from the diagram is the high level of connectivity among Graphic, Named Alternative Group, and Group Objects. Due to space constraints, the full connectivity of a few Objects (e.g., Reaction Step, Color, and Font) is not depicted. Including such relationships would add considerable complexity to an already complex system.
Perhaps surprisingly, Node and Bond, although central features in every other cheminformatics format, play a relatively minor role here. Most of the information associated with these two Objects is carried by attributes on the corresponding XML nodes.
In contrast, the objects appearing near the bottom of this diagram have relatively few associations. For example, we can see that a TLC Plate has zero or more TLC Lanes, each of which has zero or more TLC Spots. None of the child Objects in this lineage refers to its parent. An outlier is Table, which can be associated with many pages and which itself can have many pages.
Although most of the lines connecting Objects are solid, some are dashed. Solid lines represent ownership (nesting) relationships. For example, a Page is nested within a Document. In terms of XML itself, a Page node is a child of a Document node. Dashed lines represent reference relationships. CDXML Objects carry an optional id attribute. One Object references another by including its id attribute value as an attribute value. CDXML Object can reference a single Object, or a list of them, by reference.
One of the more serious limitations of the PE specification is that it fails to identify quantifier relationships between some Objects. The specification has no systematic way to report this information, opting instead to include it in within text descriptions. As a result, the quantifiers presented in the above diagram are drawn when possible from freeform descriptions found in the specification, and my own reverse engineering attempts otherwise.
Conclusion
CDXML is an important file format due to the ubiquity of ChemDraw in chemistry. Although a written specification exists, it is objectively incomplete. This article takes a first step toward filling in some of the gaps with a description of the CDXML format from a high level using ERD as a graphical tool.