The FDA Unique Ingredient Identifier (UNII)
Did you know that United States Food and Drug Administration (FDA) maintains a Substance Registration System (SRS) that aims to centralize information on every ingredient contained in domestically-marketed foods, drugs, and medical devices? A central component of this system is the Unique Ingredient Identifier (UNII). The key documentation and resources on this system are scattered and extremely difficult to locate. This article summarizes some of the more important points about the SRS/UNII initiative from the perspective of cheminformatics.
What is it?
According to the SRS Website:
The overall purpose of the joint FDA/USP Substance Registration System (SRS) is to support health information technology initiatives by generating unique ingredient identifiers (UNIIs) for substances in drugs, biologics, foods, and devices. The UNII is a non- proprietary, free, unique, unambiguous, non semantic, alphanumeric identifier based on a substance's molecular structure and/or descriptive information.
A lot of information about the SRS is contained in the FDA Substance Registration System User's Guide Version 5c. This 88-page document explains in great detail how to prepare a registration for the SRS. For example, these are the guidelines for registering substances containing azide groups:
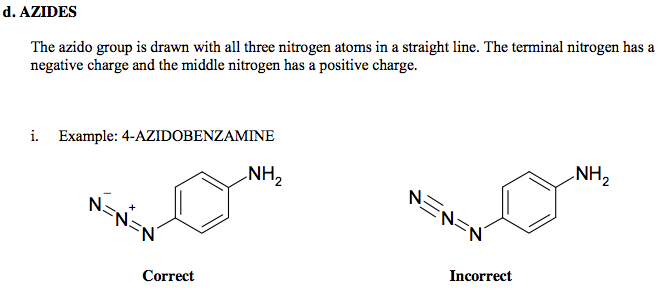
What are UNII Codes?
One of the more interesting aspects of the SRS is the structure and purpose of UNII codes. From the SRS FAQ:
A UNII is an identifier for a single defined substance. UNIIs contain 9 randomly generated alphanumeric characters with a tenth check alphanumeric character. Database managers may request the check character algorithm from fda-srs@altum.com. In order to generate a UNII, a certain amount of information must be available to unambiguously identify a given substance. The assignment of the same UNII to two materials does not indicate that these materials are equivalent in preparation method, impurity profile, or utilization, but rather only indicates that they represent the same molecular entity or elements upon which the definition is based.
All well and good, but how do we use this system to find information about substances?
Substance Lookup
The NIH offers a service that can be used to find a UNII based on substance name or vice versa:
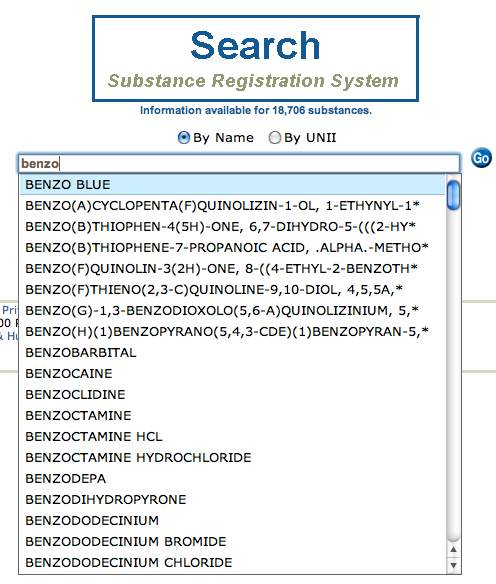
Download the UNII to Substance Name Dictionary
The FDA documentation makes clear the non-proprietary nature of UNII codes, and the copyright statement makes clear that the entire database is a public-domain work, like most works created by the federal government. So SRS and its contents can safely be repurposed and mashed up to create new services and databases - if the source files can be located.
However, locating a link to download the up-to-date contents of the FDA's UNII database from the FDA site was an exercise in frustration. By pure chance I stumbled on the UNII dictionary download page. The link can also be accessed (through a couple of hops) via the NIH's UNII lookup site.
The Excel file obtained by clicking on the download link contained the following columns:
- Preferred Substance Name
- UNII
- Substance Name
- ITIS TSM
- Molecular Formula
The file contained 61,804 entries. Although each UNII represents a single substance, multiple substance names can be associated with one UNII. In other words, this file makes it possible to disambiguate many of the most widely-used substance names. The presence or absence of the Molecular Formula field appeared to be a good indicator of whether the substance represented a discrete molecular entity (e.g., "OROTIC ACID MONOHYDRATE") or a complex mixture (e.g., "RABBIT HAIR"). The presence of an "n" in the formula suggests a polymeric material (e.g., "TRITON X 100").
It's unclear how often this download file is updated, but the date in the filename indicates that the copy I received was created on October 11, 2010 - about a week ago.
Conclusions
The FDA's SRS system, and more particularly the information it contains, can be a useful addition to any chemical database. With a few well-marked signposts it should be possible to extract useful chemical information out of free government databases using UNII as the linking mechanism. Stay tuned for more.