Looking at InChIs
InChI identifiers can be viewed both as unique molecular keys and as a language encoding molecular structure. With the right software, it is possible to decode any InChI to arrive at a human-readable molecular structure. This tutorial will show how to convert InChI identifiers into 2-D molecular renderings using open source tools.
Prerequisites
The InChI to 2-D image conversion process requires two pieces of software:
- Rino decodes InChI identifiers into molfiles. The resulting atomic coordinates are set to zero.
- RCDK assigns coordinates to the molfile produced by Rino, and renders the result.
Bring on the Code
The following Ruby code illustrates how the InChI for the pesticide fipronil (Regent) can be translated into a PNG image:
require 'rubygems'
require_gem 'rino'
require_gem 'rcdk'
require 'util'
inchi = 'InChI=1/C12H4Cl2F6N4OS/c13-5-1-4(11(15,16)17)2-6(14)8(5)24-10(22)9(7(3-21)23-24)26(25)12(18,19)20/h1-2H,22H2' #fipronil
reader = Rino::InChIReader.new
molfile1 = reader.read(inchi) # lacks 2-D atomic coordinates
molfile2 = RCDK::Util::XY.coordinate_molfile(molfile1) # has 2-D atomic coordinates
RCDK::Util::Image.molfile_to_png(molfile2, 'fipronil.png', 350, 300)
Running this code produces the image fipronil.png in your working directory:
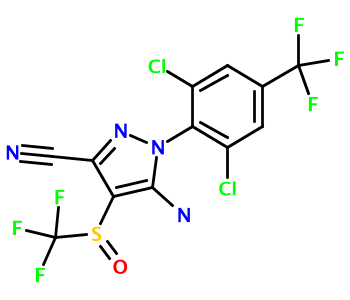
Limitations
The technique illustrated here is subject to the same limitations as the underlying software. For Rino, this means that stereochemistry is ignored. For RCDK, this means that implicit hydrogen atoms, isotopes, and charges are omitted, and that layout of macrocycles and other complex ring systems may not subjectively appear very refined.
Other Software that Does This
To my knowledge, only one other Open Source package, BKChem, is capable of rendering InChIs as described here. BKChem's underlying InChI translation and depiction software, OASA, can also be accessed online. For comparison, OASA produces the following image for for the fipronil InChI:
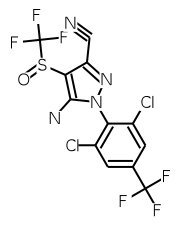
The PubChem editor can also translate and render InChIs, but no source code appears to be available. PubChem's InChI translation and rendering output for fipronil is:
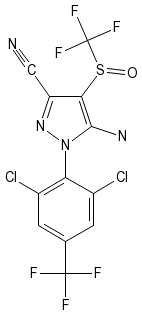
The Chemistry Development Kit, on which RCDK is based, was recently upgraded to support reading InChI identifiers. For some time, CDK has been able to generate 2-D atomic coordinates.
More information on InChI software can be found at Beda Kosata's InChI.info site.
The Final Word
Within certain limitations, it is quite feasible to programatically obtain a 2-D molecular image for any InChI identifier. Combining this capability with other chemical informatics software and services offers numerous possibilities to use InChI in innovative ways.