Hacking Molbank: Creating a Graphical Table of Contents
Molbank is an Open Access collection of single-compound articles on synthetic chemistry. Previous articles on Depth-First have highlighted Molbank's practice of including machine-readable molecular representations of its content, and its very liberal policy on mirroring and robots. In this article, we'll take advantage of both of these features to build something that was left out of Molbank: a graphical table of contents.
The Graphical Table of Contents (GTOC)
The Molbank Graphical Table of Contents (Molbank GTOC) is available online. It consists of a single Web page containing a grid of color 2-D chemical structures representing the contents of Molbank. Each structure is hyperlinked into the Molbank site itself. Clicking on the structure takes you to the complete synthetic procedure and characterization data.
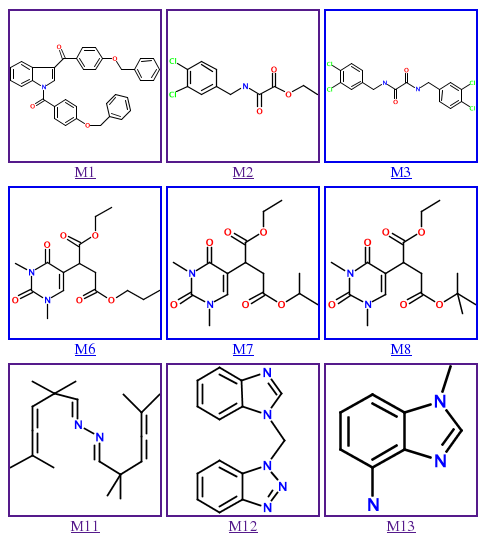
Prerequisites, Downloading, and Running
To run this project, you'll need Ruby CDK. A recent article described the small amount of system configuration required for Ruby CDK on Linux. Another article showed how to install Ruby CDK on Windows.
The complete source code for this project can be downloaded from RubyForge. A subdirectory called demo contains the pre-built final result.
After unpacking the molbank-0.1.0 archive, the demo application can be run:
cd molbank-0.0.1
ruby test.rb
Problems, We've Got Problems
Several problems were uncovered while building the Molbank GTOC. This is to be expected with any data produced "in the wild" rather than within the safety of an Ivory Tower. Here are the main categories:
- Blank Images The entry for M52 is blank. Checking the underlying molfile reveals four instances of bond stereo flags set to "6," a problem common to many of the blank images in the GTOC. According to the Molfile specification, a value of 6 indicates "Down, double bonds," whatever that means. Given that the molecules shown in M52 only have one possible stereo bond, and that the Molfile specification relies on 2-D coordinates to encode double-bond geometry, an encoding inconsistency or incorrect stereo interpretation may be the cause.
- Images Containing an "R" Atom Label Entry M53 shows an "R" group at what should be the carbonyl carbon. The underlying molfile contains several less-common entries in the properties block, a common feature of images containing "R" in the GTOC.
- Molfile not Found Entry M95 has no associated Molfile because it simply reports errata for other articles. M253-M259, on the other hand, lack molfiles because the articles were "withdrawn before publication." M347 describes a cyclodextrin for which, understandably, no molfile was provided. There are also a couple of cases in which a link to a molfile is provided, but is not available, such as M352.
- Broken Molfiles The Molfile for M162 encodes its line endings as two carriage returns and a newline, giving rise to the appearance of blank lines after data lines. This is something the Molfile specification strictly forbids. Apparently, the underlying CDK molfile reader can only handle one carriage return and a newline. Perhaps the extra return was introduced as the file was copied into and out of text editors on various operating systems in preparation for uploading it to Molbank. Another common problem was binary files being used for molfiles, such as with M402. These files don't appear to be compressed with either Zip or GZip and their nature is currently unknown.
- Bogus Molfiles For reasons I still can't understand, the Molfile for M407 encodes ethylene. So do several other Molbank molfiles. Other common dummy molfiles include toluene, benzene, and ethane.
After cataloging the problems that exist with the Molbank dataset and the software used to mine it, two interesting questions come into focus:
- What can be done to help Molbank fix the most obvious problems in their molfiles and would they accept these improvements?
- How can "real" datasets like Molbank help developers build better cheminformatics software? (a graphical Molfile Debugger Utility would come in handy…)
Clearly, the connection between Open Access, Open Source, and Open Data is very strong and runs very deep.
Behind the Scenes
The Ruby Molbank GTOC generator works by connecting to the www.mdpi.net server to get its data in real-time. Internally, the software creates a map of the Molbank website so that the molfile (and URL) for any article can be retrieved on demand. Each readable molfile is used to create a 2-D image using Ruby CDK. As a final step, the index.html page is generated, linking the 2-D images to a specific URL for a Molbank article. This file is produced with eRuby using a previously-described technique.
Conclusions
Building a Graphical Table of Contents for Molbank is not that difficult given the power of Ruby, and Molbank's forward-thinking attitude toward mirroring and robots. In working on this project, several problems were uncovered, both with Molbank's data, and the software used to mine it.
In some ways, the software described here and its output are less interesting than the larger questions they raise:
- How do scientific journals best serve not only their readers, but developers who want to provide new ways to use the journal?
- How far does copyright extend in scientific publications? For example, are molfiles copyrightable? If so, at what level of detail are they not? If atom coordinates or some other kind of non-essential information is left out, does that change anything?
- In what other practical ways could the connection between Open Source, Open Data, and Open Access be explored?
These and many related questions are waiting just around the corner. As Open Access becomes more viable, both technically and commercially, look to Open Source and Open Data to provide the synergies that will unlock its true potential.